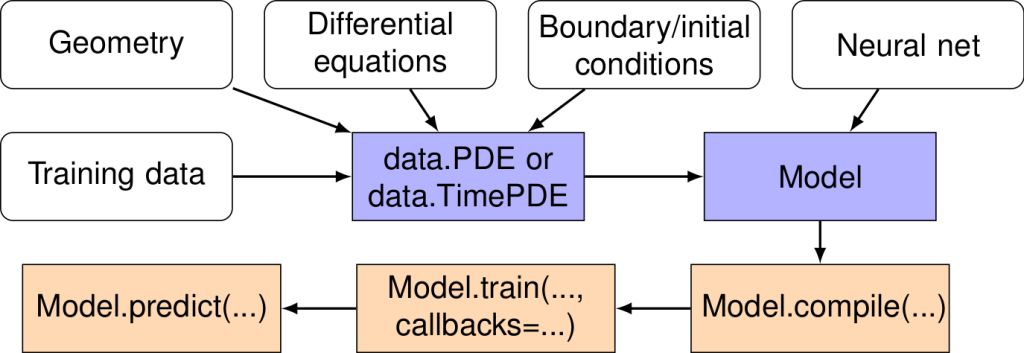
DeepXDE
- A library for scientific machine learning and physics-informed learning.
- >900,000 downloads, >2,900 GitHub Stars, >75 Contributors. Used in hundreds of papers published by a diverse range of scientists from >200 universities, national labs, and industry.
OpenRBC
- A coarse-grained molecular dynamics code for simulating entire human red blood cells at the protein resolution. The code outperforms a legacy solver by 10 times in time-to-solution through the use of a novel adaptive spatial searching algorithm to accelerate the computation of short-range pairwise interactions in an extremely sparse 3D space.
- Third Prize, IBM OpenPOWER Developer Challenge contest, 2016.
